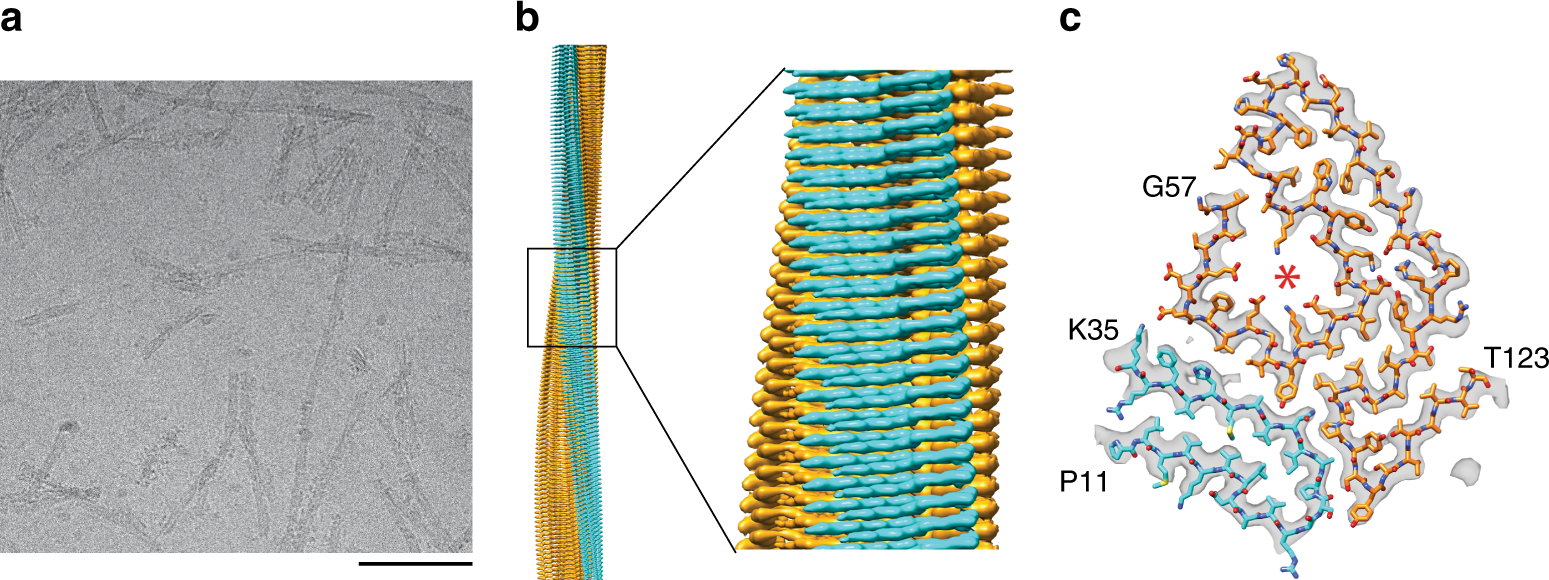
Cryo-EM structure of a transthyretin-derived amyloid fibril from a patient with hereditary ATTR amyloidosis | Nature Communications

Using extensional flow to reveal diverse aggregation landscapes for three IgG1 molecules - Willis - 2018 - Biotechnology and Bioengineering - Wiley Online Library

Example of AGGRESCAN output. The red line represents the aggregation... | Download Scientific Diagram
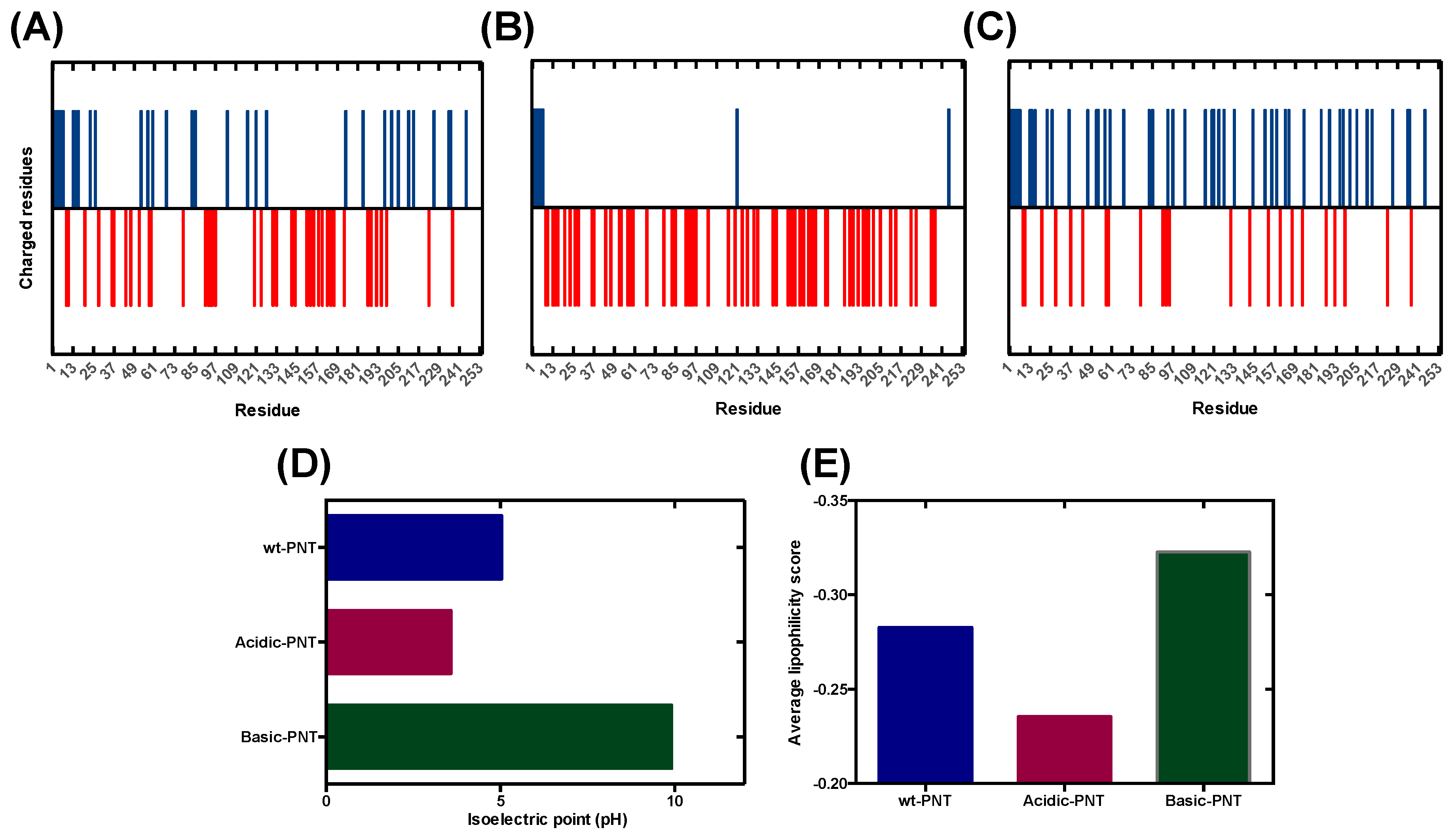
Cells | Free Full-Text | pH-Dependent Aggregation in Intrinsically Disordered Proteins Is Determined by Charge and Lipophilicity | HTML

Computational prediction of protein aggregation: Advances in proteomics, conformation-specific algorithms and biotechnological applications | Semantic Scholar

AGGRESCAN: a server for the prediction and evaluation of "hot spots" of aggregation in polypeptides | BMC Bioinformatics | Full Text
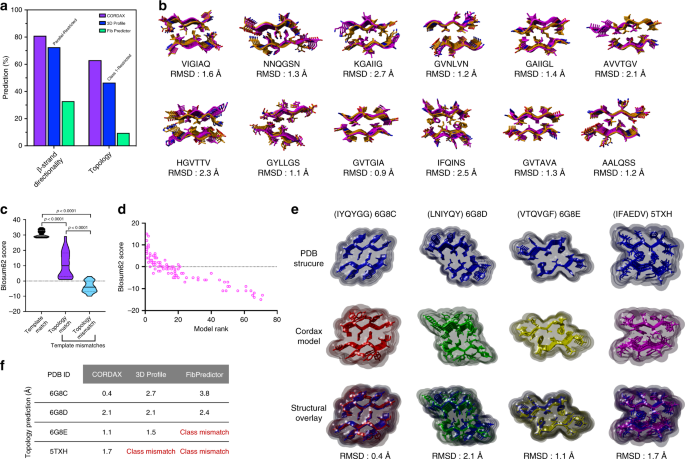
Structure-based machine-guided mapping of amyloid sequence space reveals uncharted sequence clusters with higher solubilities | Nature Communications

AGGRESCAN: a server for the prediction and evaluation of "hot spots" of aggregation in polypeptides | BMC Bioinformatics | Full Text

AggScore: Prediction of aggregation‐prone regions in proteins based on the distribution of surface patches - Sankar - 2018 - Proteins: Structure, Function, and Bioinformatics - Wiley Online Library

Example of AGGRESCAN output. The red line represents the aggregation... | Download Scientific Diagram

PDF) AGGRESCAN: a server for the prediction and evaluation of "hot spots" of aggregation in polypeptides | Xavier Daura - Academia.edu
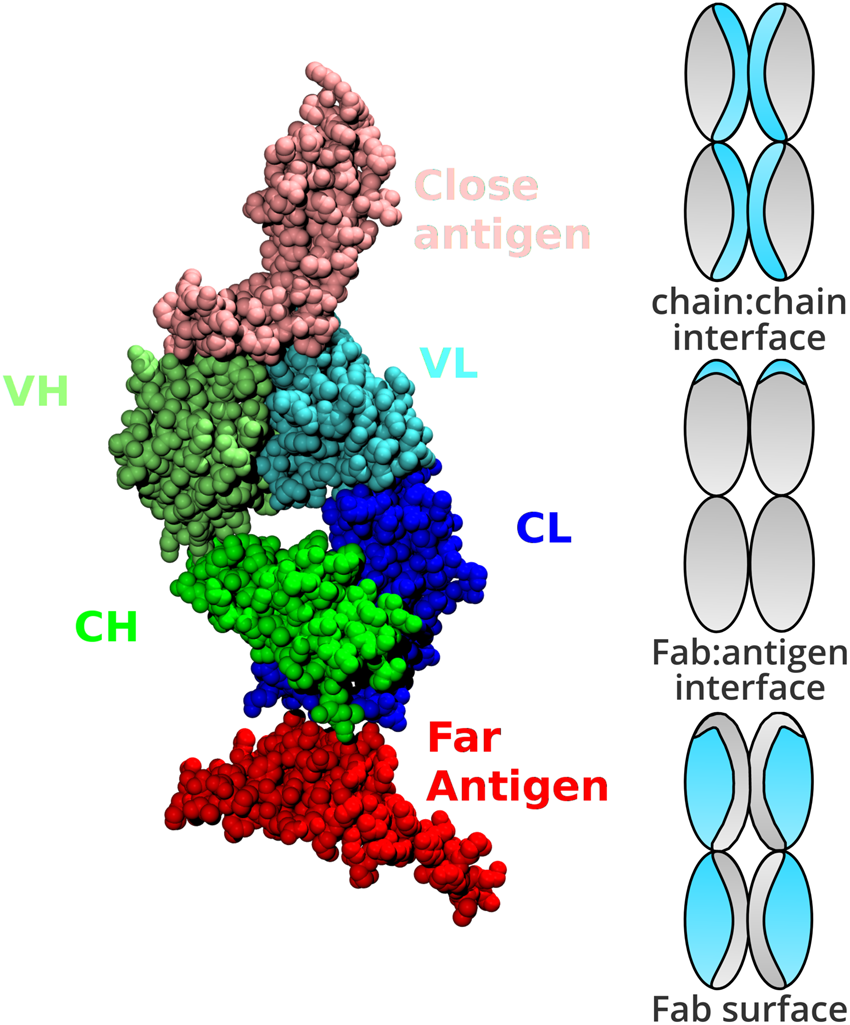
Web-based display of protein surface and pH-dependent properties for assessing the developability of biotherapeutics | Scientific Reports

Aggregation-prone regions. (a) Aggregation-prone regions predicted by... | Download Scientific Diagram

AggScore: Prediction of aggregation‐prone regions in proteins based on the distribution of surface patches - Sankar - 2018 - Proteins: Structure, Function, and Bioinformatics - Wiley Online Library
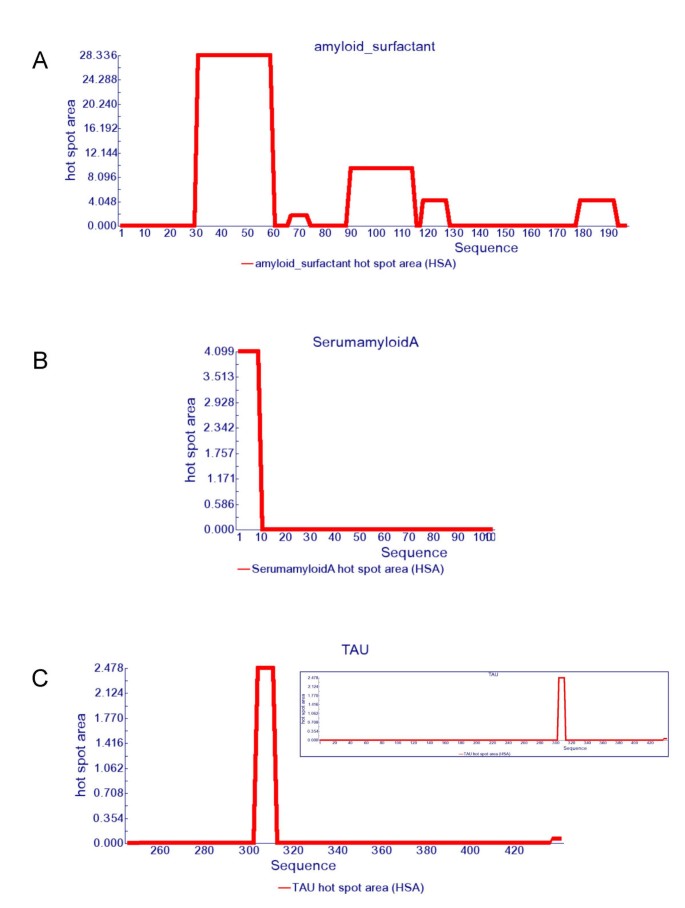
AGGRESCAN: a server for the prediction and evaluation of "hot spots" of aggregation in polypeptides | BMC Bioinformatics | Full Text
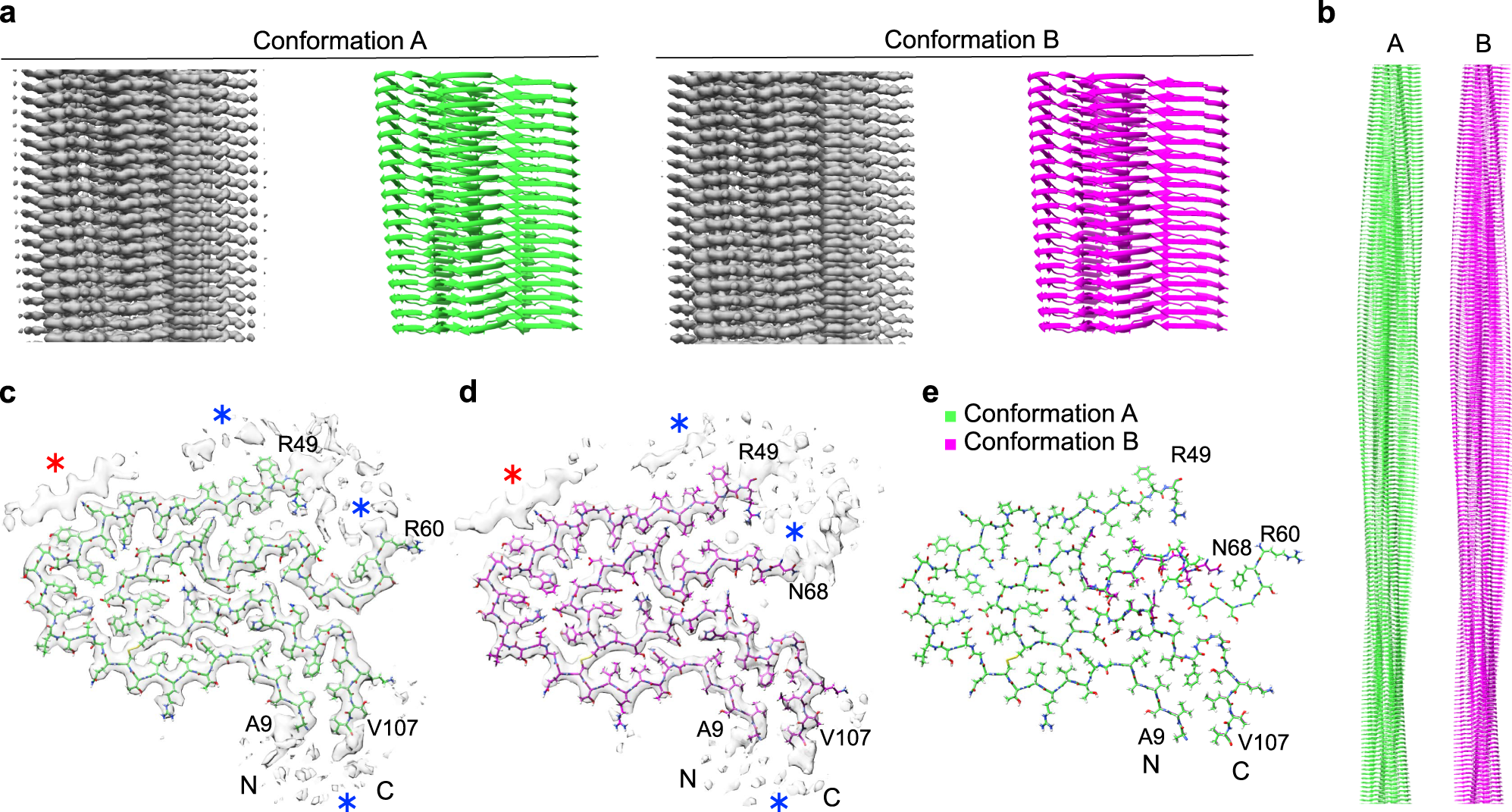
Cryo-EM reveals structural breaks in a patient-derived amyloid fibril from systemic AL amyloidosis | Nature Communications
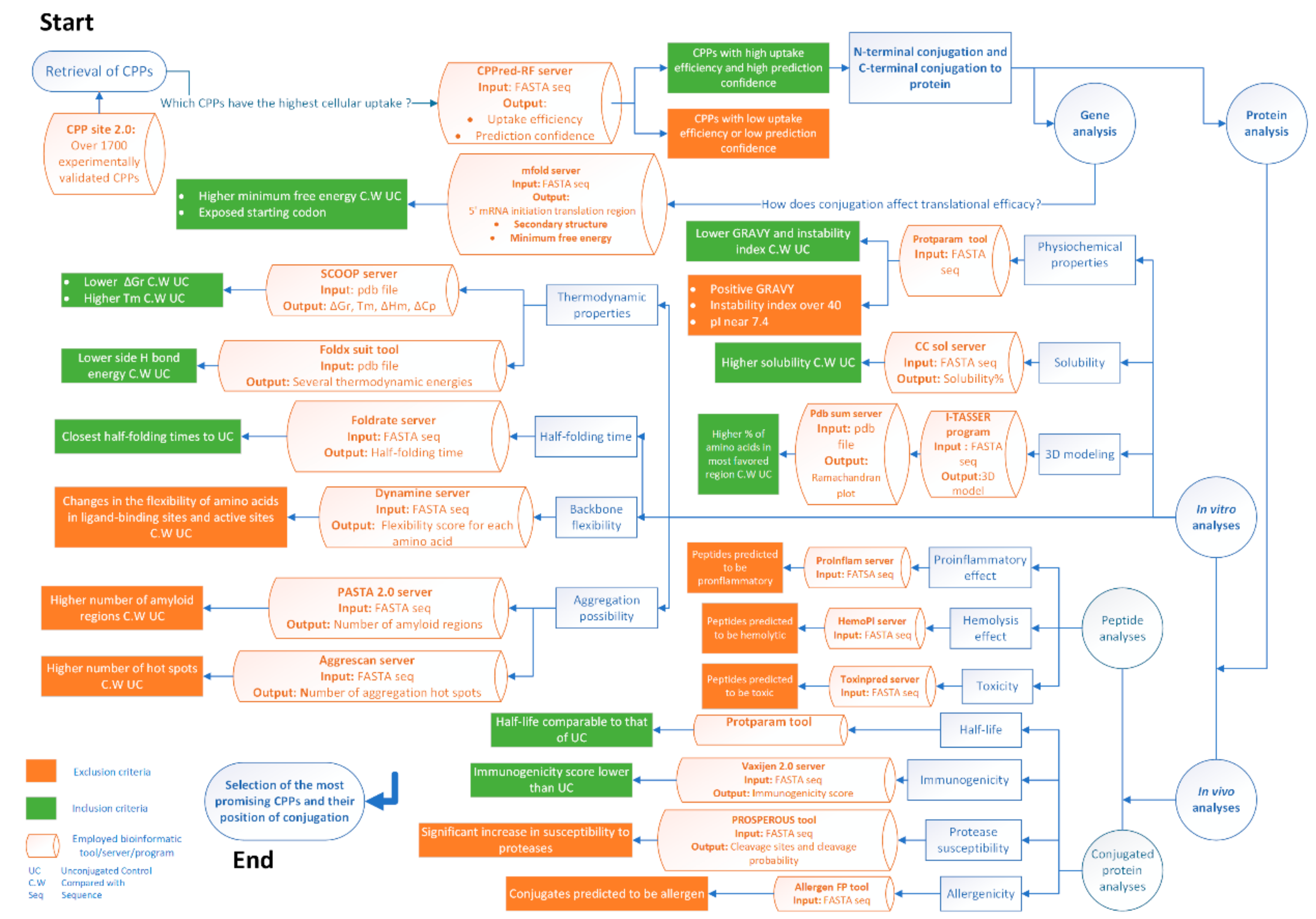
Molecules | Free Full-Text | Considerations on the Rational Design of Covalently Conjugated Cell-Penetrating Peptides (CPPs) for Intracellular Delivery of Proteins: A Guide to CPP Selection Using Glucarpidase as the Model Cargo
![PDF] AGGRESCAN3D (A3D): server for prediction of aggregation properties of protein structures | Semantic Scholar PDF] AGGRESCAN3D (A3D): server for prediction of aggregation properties of protein structures | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/bc8d0acd077eed82b977fbf9d706415235ef1fe6/4-Figure1-1.png)
PDF] AGGRESCAN3D (A3D): server for prediction of aggregation properties of protein structures | Semantic Scholar
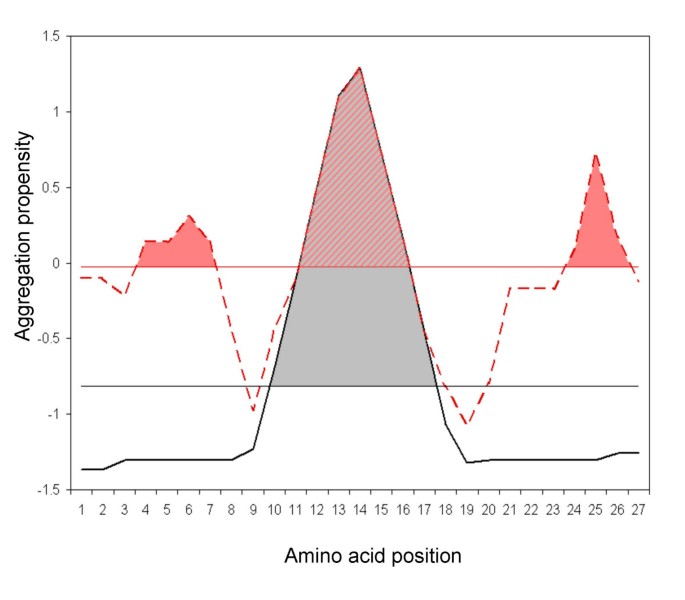
AGGRESCAN: a server for the prediction and evaluation of "hot spots" of aggregation in polypeptides | BMC Bioinformatics | Full Text

![PDF] Guidelines to reach high-quality purified recombinant proteins | Semantic Scholar PDF] Guidelines to reach high-quality purified recombinant proteins | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/1906620dcc5549d1883d5dedeef415889614d0a7/4-Table1-1.png)



